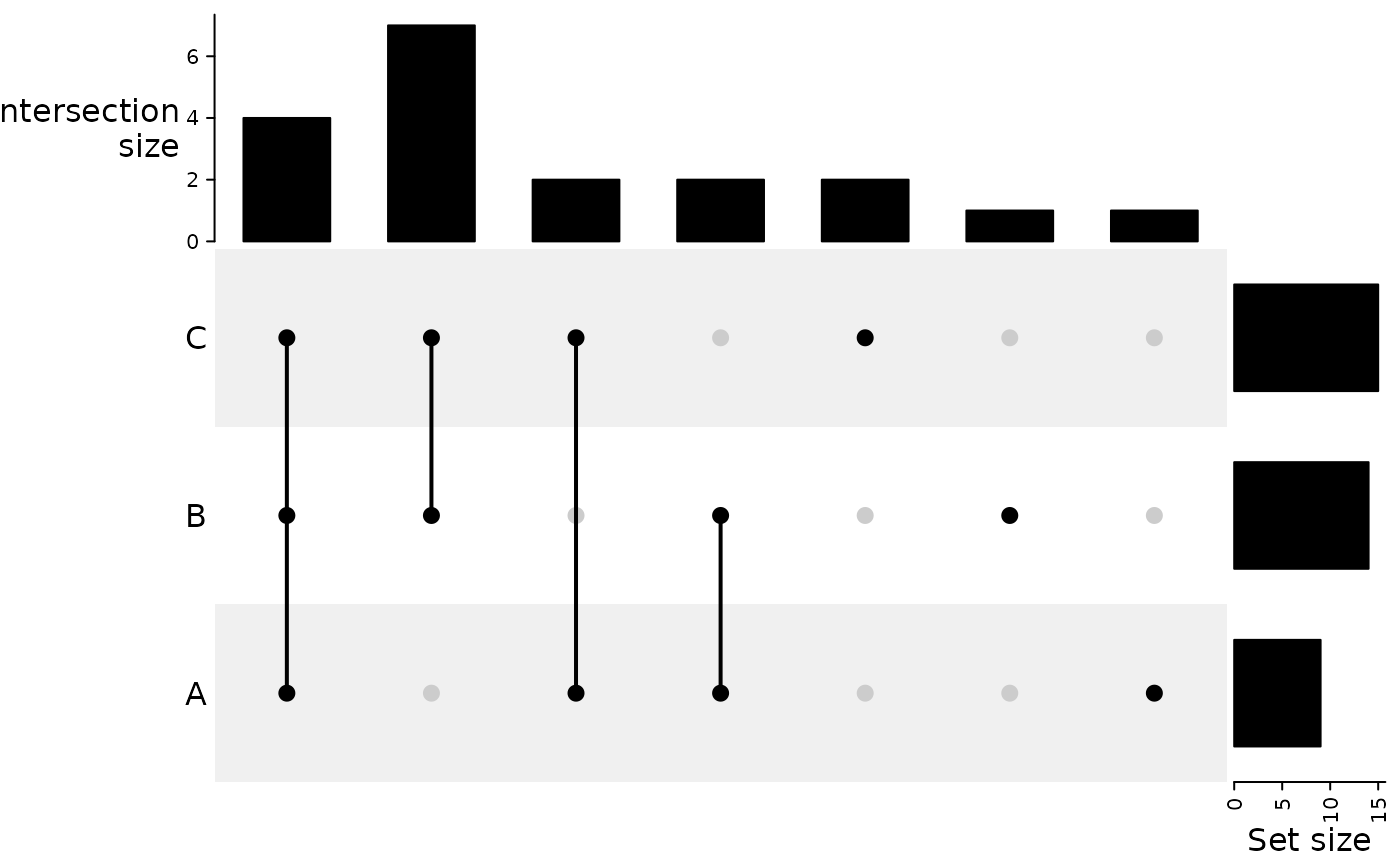
Prepares sets of regions for UpSet overlap representation.
Usage
regionsToUpset(
x,
reference = c("reduce", "disjoin"),
returnList = FALSE,
ignore.strand = FALSE,
maxgap = -1L,
minoverlap = 0L,
...
)Arguments
- x
A named list of genomic ranges (or paths to bed files)
- reference
The method for creating the reference windows ('reduce' or 'disjoin'). Alternatively, a `GRanges` object of reference windows.
- returnList
Logical; whether to return the list of regions instead of plotting.
- ignore.strand
Logical; whether to ignore strand for overlaps (default FALSE).
- maxgap
Max gap between regions to consider an overlap.
- minoverlap
Minimum number of overlapping bases.
- ...
Further arguments specifying how the overlaps are done, passed to
findOverlaps-methods).
Value
A data.frame of set inclusions which can be directly input to
make_comb_mat, and then
UpSet.